GenCore operates a combined total of five different next generation sequencing platforms across its two locations.
GenCore is an Illumina Certified Service Provider (CSPro).
The core technical staff has extensive expertise in experimental design, library construction, and operation of next generation sequencing equipment.
The core bioinformatics staff provide raw data processing and are available for
consultation for a variety of downstream analyses
New York
GenCore manages sequencing operations using TuboWeb, our in-house LIMS system.
In order to sequence, researchers must input library metadata into TuboWeb. This includes information such as pool name, library names, barcodes, and final pool concentration.
The steps for sequencing at GenCore are outlined below.
Printable steps can be found here.
Add multiple libraries using the CSV template upload
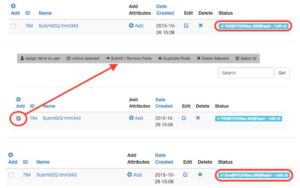
Assign one or multiple libraries to a pool
CSV upload will automatically create a pool
The GenCore manager will send an email confirmation that includes the submission deadline and scheduled run date
Quantify libraries using the KAPA protocol within 5 days of your scheduled run date
Do not freeze/thaw the pool after doing KAPA
Add final concentration and volume of pool by clicking on the pool name and then “Add Vol & Conc”
Place on the “Sequencing Submissions” rack in 4°C fridge
Provide the TapeStation results for your final pool by replying to the confirmation email
Abu Dhabi
The Human Genome Project was completed in ten years and cost three billion dollars. Due to recent advances in high-throughput sequencing technology, an entire human genome can now be sequenced in a few days for one millionth of the original cost. The Sequencing Core Technology Platform is used by researchers in genomics and systems biology to rapidly collect large amounts of data on DNA and RNA sequences from humans, vertebrate and invertebrate animals, plants, and microbes in order to characterize their genomes (DNA), transcriptomes (RNA), and genetic diversity in populations (sequence variants).
Applications include the analysis of genomes, gene expression patterns, regulatory interactions, epigenetics, and how sequences vary within populations and between species. Microbes: Applications include the analysis of genomes, gene expression patterns, regulatory interactions, epigenetics, and how sequences vary within populations and between species. Other applications include investigating interactions between proteins and nucleic acids (e.g., using ChIP-seq or PAR-CLIP) and analyzing samples collected from the environment (metagenomics).
Data analysis for the Sequencing CTP is supported by NYUAD-CGSB’s Bioinformatics Core.