All library documentation, metadata, and associated files are encoded through the Genomics Core LIMS: TuboWeb. The sequencing queue is also managed through TuboWeb.
NYU TuboWeb
- TuboWeb Video Tutorial
- Note to First Time Users: After initial login, one must request to be added to the lab group. This can be completed by your lab administrator or the GenCore Bioinformatics Specialist.
My Dashboard…
gives an overview of current status of libraries, samples in the sequencing queue, and planned runs.
Projects…
allows the creation of a Project to sort data from multiple pools and libraries into one category. This is optional.
My Group…
allows for other researchers to be assigned to an individuals group for metadata sharing.
Libraries…
is where all the metadata for each sequencing library is encoded. All sequencing libraries in the system can be viewed as well as their validation status. If there is missing metadata, a flag will alert to what the missing or incorrect information is. Here is where protocols, QC files, and etc. can be uploaded and associated with a specific library.
Files…
allows for a bulk upload of files (protocols, gel images, BioA traces, and etc.) that can be then associated with sequencing libraries.
Import csv…
allows for the quick and easy import of all metadata. The template can be found for download on this tab. The template mirrors that of previous GenCore submission forms and helps to facilitate encoding of large sequencing runs.
All Pools…
is where all sequencing pools can be encoded and all generated sequencing pools are be viewable. Here validation flags can be viewed and what the pool status is, ie submitted, in progress, or sequenced. Important specifications for sequencing can be noted here, ie whether custom sequencing primers are required, if the pool contains Nextera libraries, and any addition comments.
Queued Pools…
shows the sequencing pools which have been submitted to the sequencing queue and are ready to be scheduled for a sequencing run.
Pools in Progress..
shows the sequencing pools that are on a scheduled, confirmed, and current sequencing run.
Sequenced…
shows the sequencing pools that have been completed and sequenced.
Barcodes…
allows for the encoding of new barcode sets as well as any home-made/custom barcode sets. These can then be associated with libraries and saved for future use. Most standard/commercially available barcode sets are encoded for convenience.
Chartfields…
is where chartfields (NYU account codes) can be added. A chartfield is required for submission of a sequencing pool and is used for GenCore invoicing.
My Runs..
is where a list of all completed sequencing runs for an individual can be viewed. This is also where quality metrics and the location of the sequencing data can be found.
How to Enter the Sequencing Queue:
To enter the sequencing queue and be scheduled on a sequencing run, one MUST encode library and pool metadata into TuboWeb. The pool metadata will be validated and if passes validation, a status of R2Q (Ready to Queue) will be given to the pool. One may then submit this pool to the GenCore sequencing queue. If this is successful, the pool is given a Q’ed (queued) status to show that it has been successfully added to the sequencing queue and the GenCore can add it to a sequencing run (pool will then be given the In Progress Status).
Note: If one wishes to run the same physical pool in multiple lanes of a flowcell, the pool must be duplicated in TuboWeb (one can use the “+Duplicate Pools” button). All duplicates must also be submitted to the queue to notify GenCore of how many lanes are needed for sequencing. One physical pool may be submitted given enough volume to accommodate multiple lanes. Contact GenCore staff if there is any uncertainty.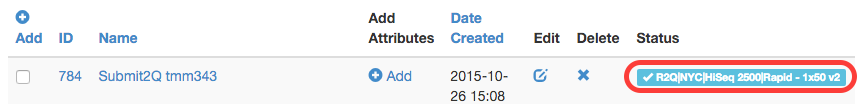
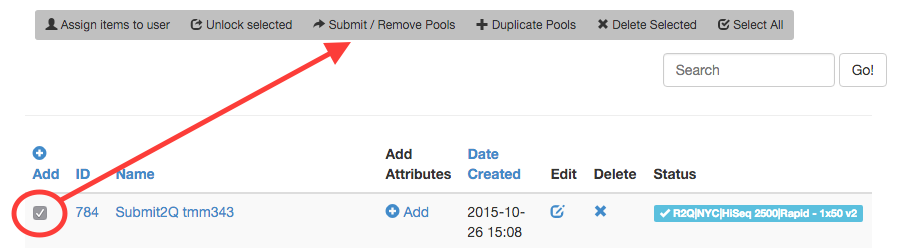
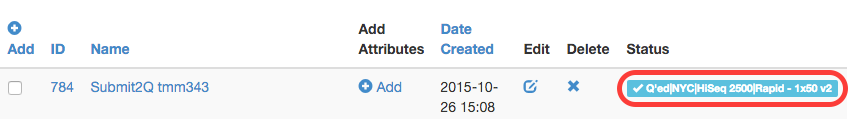
GenCore will schedule sequencing runs solely on what is submitted to the queue via TuboWeb. The queue can be viewed on the dashboard in TuboWeb and lanes available can be seen on potential sequencing runs.
When a sequencing run has been filled, one will receive a run confirmation e-mail from GenCore outlining the submission date and final steps to be taken. Do not submit physical samples to the GenCore until this e-mail has been received. Also the final required qPCR can be completed after samples have been queue’d in the system and once a run confirmation e-mail is sent. (TapeStation, BioA, Qubit should all be done to ensure successful library prep). These values can be added after queuing using the button depicted below: